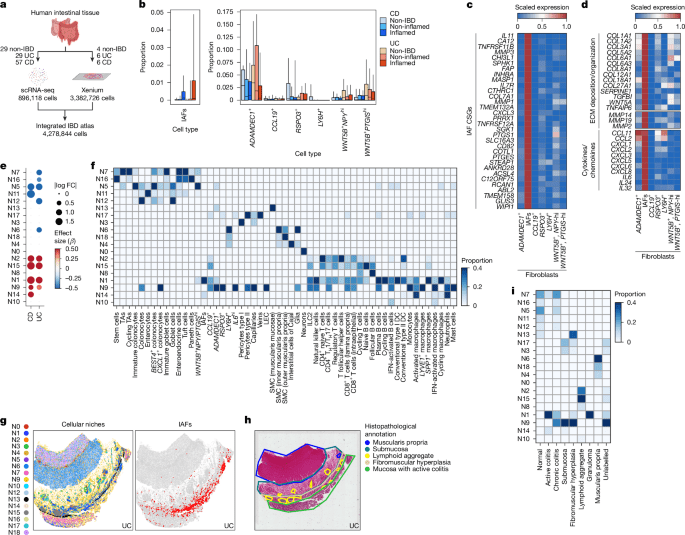
Generating the integrated IBD scRNA-seq atlas
Previously published scRNA-seq datasets on CD and UC were used for integrative analysis2,9,11,13. The dataset of Smillie et al.2 was generated with colonic tissue biopsies obtained from 12 non-IBD controls and 18 patients with UC (from both inflamed and non-inflamed regions). Libraries were prepared for single-cell profiling by fractionating the tissues into epithelial and lamina propria fractions before downstream processing. The Kong et al.13 dataset was generated from 13 non-IBD controls and 46 patients with CD and consisted of biopsies obtained from inflamed regions of 17 participants and non-inflamed regions of 43 patients with disease. Libraries for single-cell profiling were prepared in a mixed manner, with some samples separated into epithelial and lamina propria fractions and some not separated. Biopsies were obtained from three segments of the gastrointestinal tract: small bowel, terminal ileum and colon.
The Friedrich et al.11 dataset was generated by fractionation and sorting of EPCAM− and CD45− cells from colonic biopsies obtained from 4 non-IBD controls and 11 patients with UC, comprising 7 inflamed region biopsies and 4 non-inflamed region biopsies. The Martin et al.9 dataset consisted of separated lamina propria fractions of paired involved and non-involved region biopsies from 11 patients with CD. The combined dataset before downstream quality control filtering comprised 1,143,316 cells from 115 patients. Data were analysed using the scanpy implementation54.
scRNA-seq analysis and cell type identification
Gene expression normalization was performed on the combined scRNA-seq dataset to account for differences in sequencing depth across cells. Unique molecular identifier (UMI) counts were normalized by the total UMI count per cell, and the total count for each cell was set to 10,000 transcripts per cell. After natural logarithm conversion and scaling of the gene expression matrix, the top 2,000 most highly variable genes were selected for a first round of dimensionality reduction. Thereafter, batch correction was performed on each individual patient sample using harmony55, followed by neighbourhood clustering and uniform manifold approximation and projection (UMAP) embedding of the single cells56. On the basis of expression of known markers of epithelial (KRT8, EPCAM), stromal (PDGFRA, PECAM1, ACTA2, S100B, RGS5) and immune (CD79A, MZB1, CD3D, TRAC, C1QA, TPSAB) cell populations, the clusters were then subdivided into these three main compartments for subsequent rounds of clustering and analysis.
For compartment-specific analyses, gene expression normalization was performed by excluding genes with higher UMI counts (more than 5% of the total UMI count per cell) to minimize the contribution of highly expressed genes to the normalization.
In the stromal compartment dataset, only genes expressed in more than five cells were considered for further integrative analysis. Dimensionality reduction and batch correction was performed by adjusting for the following covariates: 10x Genomics Single Cell Gene Expression Solution chemistry (v.1, v.2 or v.3), patient, study and tissue site (small bowel, ileum or colon). The top 45 adjusted principal components were considered for neighbourhood clustering using the Leiden algorithm57 and visualized using UMAP embedding. This was followed by post hoc analysis of identified clusters to remove poor-quality cells; that is, clusters with low UMI counts or high mitochondrial gene fraction and those expressing lineage markers of non-stromal cells were removed as doublet cells. Wilcoxon rank-sum test was performed to define the markers specific to individual clusters and annotate each cluster.
Analyses for the epithelial and immune compartments followed a similar workflow to that described above, except that further rounds of iterative clustering were performed after removal of doublet and low-quality cells.
Generation of a Xenium-based spatial transcriptomics dataset
Human sample collection
Sixteen patients diagnosed with UC, CD or diverticulitis who were recruited into the Prospective Registry in IBD Study at MGH (PRISM) study at Massachusetts General Hospital (MGH) participated in this study. Informed consent was obtained from all patients in accordance with the protocol approved by the institutional review board (IRB; 2004P001067). The study protocol complied with all relevant ethical regulations. Samples from patients with diverticulitis were pathologist-confirmed histologically normal. Excess tissues from clinically warranted surgical resections were collected for research purposes. IRB-approved secondary use protocol 2020P001262 allowed use of these tissues in research at the Broad Institute of MIT and Harvard.
Human colon sample processing
Sample preparation for Xenium profiling
Human colon FFPE blocks were sectioned at 5 µm thickness on a microtome (Leica HistoCore AUTOCUTl) and placed on Xenium glass slides (10x Genomics) following slide equilibration at room temperature for at least 30 min. Slides were baked at 42 °C for 3 h and thereafter stored in a desiccator overnight. Tissues were baked at 60 °C for 2 h, deparaffinized in xylene (MilliporeSigma, 214736-4L), ethanol rehydrated and decross-linked according to the manufacturer’s protocol (10x Genomics, demonstrated protocol CG000580 rev. C). Protocol CG000582, rev. F (10x Genomics) was used for probe hybridization, ligation and amplification for eight of the samples, whereas user guide CG000749 rev. B (10x Genomics) with further cell segmentation staining was used for the other eight human samples. A customized 480-plex gene probe panel (probe ID: HC3GPZ) was designed and used for all human samples. The probes were hybridized on to the tissue for 19 h at 50 °C.
When used, the slides were incubated with cell segmentation markers for 17 h and 30 min at 4 °C. Buffers were prepared for the Xenium Analyzer instrument (software v.3.1.0.0) according to the manufacturer’s protocol for the Xenium v1 workflow (10x Genomics, user guide CG000584 rev. G). Selected regions of interest covering the full tissue of each sample were imaged, and spatially localized transcriptional data were collected.
Mouse colon tissue blocks embedded in optimal cutting temperature (OCT, Tissue-Tek) were cryosectioned at 10 µm thickness in a cryostat (Leica, CM1950) at −20 °C. Tissue sections were placed on Xenium glass slides (10x Genomics) and stored in a slide mailer at −80 °C for 4 days. Tissues were fixed in 4% paraformaldehyde (Thermo Fisher Scientific, BP531-25) and permeabilized according to the manufacturer’s protocol (10x Genomics, demonstrated protocol CG000581 rev. D). Tissue slides were further processed following the 10x Genomics user guide (CG000749 rev. B). A customized 480-plex gene probe panel (ID: JUR2CE) was used, with hybridization for 21 h at 50 °C. Tissues were stained with cell segmentation markers for 17 h and 30 min at 4 °C. Buffers for the Xenium Analyzer instrument (software v.3.2.0.1) were prepared following the Xenium v1 workflow (10x Genomics, user guide CG000584 rev. G). Slides were imaged, and transcriptional data were collected using a Xenium Analyzer instrument.
Post-Xenium tissue slides were incubated for 10 min with 10 mM sodium hydrosulfite (Sigma-Aldrich) to remove the tissue quencher (10x Genomics, demonstrated protocol CG000613 rev. B) and H&E stained using the automatic H&E stainer at the MIT Histology Facility. Glass slides were imaged on a Zeiss Axio microscope with a Metafer slide-scanning platform using a ×10 objective.
Analysis of human spatial transcriptomics dataset
Samples were processed in two batches, with and without the cell segmentation staining kit. For tissues processed without the cell segmentation staining kit, we used a nucleus segmentation mask to restrict the counting of transcripts per cell located in its nucleus. In total, a gene expression matrix of 4,477,548 cells × 480 genes was obtained from 16 human tissues and analysed using a similar workflow to that described for the scRNA-seq dataset (see Supplementary Data 3 for the panel gene list). The analysis was implemented in Scanpy. In brief, quality control was performed to remove poor-quality cells with total RNA counts less than 10. Count normalization and log transformation were followed by regression of the effect of total RNA per cell and scaling. Principal component analysis was conducted to generate the top 50 principal components.
Batch correction was then performed using the harmony package, with variables patient and batch as covariates After inspection of the elbow plot, we used the top 30 eigenvectors to construct a k-nearest neighbour graph with n_neighbors = 15. Subsequently, the Leiden clustering algorithm was applied (resolution = 0.5) to cluster the spots, and the resulting clusters were visualized with the UMAP embedding algorithm. On the basis of the expression of known markers of epithelial cells (KRT8, EPCAM), fibroblasts (PDGFRA, C1S), stromal cells (PECAM1, ACTA2, DES, S100B, RGS5, CD9, ANO1), lymphocyte populations (CD79A, MZB1, CD3D, TRAC), and myeloid cells (C1QB, CD83, AIF1, TPSAB, CD36, FCGR3B), the clusters were then subdivided into five main compartments for subsequent iterative rounds of clustering and analysis.
In contrast to the scRNA-seq analysis workflow, we further subdivided the stromal cells into fibroblast and non-fibroblast compartments, and likewise the immune cells into lymphocytes and myeloid cells for resolution of cell type heterogeneity.
Compartment-specific analysis and clustering was implemented using a similar workflow to that described above by setting Leiden algorithm parameter resolution = 1. Clustering was followed by post hoc analysis of identified clusters to remove poor-quality cells, that is, clusters with low total RNA counts or those expressing lineage markers from other compartments. Wilcoxon rank-sum test was performed to define the markers specific to individual clusters and annotate them. Altogether, we identified 56 unique cell types across the different compartments in the spatial atlas.
Harmonizing single-cell and spatial atlases
Analysis of the scRNA-seq IBD atlas (scIBD) revealed 51 distinct cell types, whereas 56 distinct cell types were found for the spatial IBD atlas (spIBD). Broadly, most of the cell types identified in the scIBD were also identified in the spIBD, which also recovered cell types that are known to be located deeper in the muscularis propria, that is, interstitial cells of Cajal, neurons, and smooth muscle cells located in the inner circular and outer longitudinal layer of the muscularis proporia. As the scRNA-seq was based on biopsies, these cell types were not captured. Likewise, monocytes, follicular B cells and IL6hi fibroblasts were identified only in the spIBD. On the other hand, colonic plasma B cells, naive CD8 T cells, and M cells (microfold) were captured only in the scIBD.
Moreover, we observed some cell types and states that were captured in the scIBD but could not be distinguished in the spIBD. For instance, we recovered the T helper 1 (TH1) and T helper 17 (TH17) polarized states of differentiated CD4 T cells in the scIBD, but the heterogeneity between these cells could not be resolved in the spIBD; hence, they were annotated as the same cluster (CD4 TH1/TH17). Naive and memory B cells were recovered in the scIBD, but their heterogeneity could not be distinguished in spIBD. Likewise, capillaries and arteries were distinct in the scIBD but grouped into the same cluster in spIBD. WNT5B+, NPYhi and PTGIShi cells were identified in the scIBD but could not be distinguished in the spIBD. Conversely, SPP1+ macrophages were identified in the spIBD, whereas C1Qhi and MHCIIhi macrophages could not be resolved.
For the ligand–receptor analysis (described below), we focused on the common set of cell types between the two atlases and, wherever possible, used the closest cell type as a proxy when the cell types could not be identified in the spIBD.
Cellular niche analysis on the basis of spatial transcriptomics
We evaluated the composition of cell types in the proximal neighbourhoods of each cell to identify the cellular niches (microenvironments) shared across multiple samples. For every cell in our spatial atlas, we counted the number of distinct cell types present within a radius of 30 μm, thereby generating a frequency matrix of 3,382,726 microenvironments by 56 cell types. The median number of total cells sampled within the 30-μm radius was 14 cells (range: 1–113). Next, we implemented a k-means clustering approach using the scikit-learn (v.0.22) package and evaluated the stability of clusters across a range of k values (5–35) using the silhouette score. We obtained the highest score at k = 19, suggesting the existence of 19 distinct niches (N0–N18) in our dataset.
Niche and cell type enrichment analysis
Chi-squared test of independence was performed to evaluate the enrichment of cell types in a given niche. A contingency table summarizing the count distribution of each cell type across niches in the overall spatial atlas was generated, and the chi-squared statistic (χ2) was computed using observed and expected frequencies, assuming independence between variables. Degrees of freedom were calculated as follows: d.f. = (N − 1) × (C − 1), where N is the number of distinct niches, and C is the number of cell types. P values for cell type enrichment in each niche were obtained using standardized residuals and corrected for multiple testing using the Bonferroni method. P < 0.05 was considered to indicate statistical significance.
Cell type composition analysis in single-cell and spatial atlases
For the spatial atlas, we performed the analyses at the patient level. Specifically, we first visualized the relative abundance of each niche across the different conditions, and identified niche N3 as the reference niche, as it did not change between conditions. scCODA was then implemented by adding a pseudocount of 0.001, using the non-IBD controls as reference. Niches were considered to have significantly changed in abundance if FDR < 20%. β values and log fold change in abundance (for niches with credible effects) were visualized.
Differential expression analysis on scRNA-seq atlas
Pseudobulk expression for each gene was first computed for each cell type in every patient and then analysed using the decoupler and pydeseq2 packages. Differential expression was evaluated using the respective controls for UC inflamed or CD inflamed participants, including 10x chemistry as a covariate in the model. In the UC inflamed group, genes with FDR-adjusted P < 0.05 were considered to show statistically significant differential expression; for the CD inflamed group, this threshold was reduced to FDR-adjusted P < 0.1, because fewer genes showed differential regulation among these patients.
Inference of cell communication network on the basis of spatial constraints
Intercellular cross-talk was analysed with the cellphonedb (v.5.0) package using the DEG analysis method. The scIBD was used as the reference, and separate analyses were performed for patients with UC and those with CD. First, differentially upregulated genes passing the significance threshold in inflamed tissues compared with control tissues, as described in the ‘Differential expression analysis on scRNA-seq atlas’ section, were used as the input for each cell type. As IAFs were predominantly observed in inflamed tissues, the differential expression analysis did not yield significantly upregulated genes in IAFs because of low numbers among controls. Hence, to obtain IAF upregulated genes, we performed differential expression at the pseudobulk level between IAFs and non-IAF fibroblast populations and appended the results to the input table. The threshold parameter in cellphonedb was modified to include genes with greater than 5% expression in a given cell type for scoring of interactions.
To infer spatially constrained ligand–receptor communication scores, we used the list mapping each niche to significantly enriched cell types, as described in the ‘Niche and cell type enrichment analysis’ section, as input. As some cell types identified in the scIBD could not be identified in the spIBD, as described in the ‘Harmonizing single-cell and spatial atlases’ section, we used the niche membership of the closest cell type as a proxy.
Specifically, CD4 TH1 and TH17 cells in the scIBD were assumed to be members of the same niches as CD4 TH1/TH17 cells in the spatial atlas, WNT5B+NPYhi and WNT5B+PTGIShi were considered to be the same as WNT5B+, capillaries and arteries were assumed to be the same as capillaries and arterioles, naive and memory B cells were assumed to be the same as naive B cells in the spatial atlas, and C1Qhi macrophages were assumed to localize to the same niches as LYVE1hi macrophages.
The total sum of significant ligand–receptor interaction scores were counted between a given sender–receiver pair of cell types and used to derive an adjacency matrix for network analysis. The igraph package was used to plot a directed and weighted network graph on the adjacency matrix, with edge thickness representing the sum total interaction score. Stromal compartment cells were removed from the network graph to highlight communication with IAFs and non-stromal cells.
Histopathology annotations
H&E-stained sections of each tissue were used to manually identify structures pertaining to various anatomical and pathological regions. Using Xenium Explorer (10x Genomics), we aligned H&E images to the slide, selected the regions manually and extracted cell barcodes. We ascertained the following anatomical regions: the normal mucosa, submucosa and muscularis propria; and the following pathological regions: mucosa with predominantly active colitis, mucosa with predominantly chronic colitis, fibromuscular hyperplasia, lymphoid aggregates and granuloma. Each tissue was also qualitatively graded for stage of fibrosis, ulceration and severity of inflammation.
Identification of putative driver ligands of IAF gene program
First, we identified transcription factors (TFs) enriched in IAFs on the basis of their expression levels. Cluster-specific enrichment analysis in the stromal compartment atlas was performed using Wilcoxon rank-sum test to identify genes enriched in IAFs. The list was then subset to the list of known TFs to yield 23 putative mediators. In parallel, we implemented DoRothEA to infer the activity of known TFs59,60. Then, we identified the enriched TFs in IAFs using the Wilcoxon test, considering all non-IAF stromal cells as the background. We selected TFs with a mean activity difference greater than 0.75 and P < 0.01, yielding a list of 34 putative TFs. The final list consisted of 55 unique TFs that putatively orchestrated the IAF program.
Next, we leveraged the ligand–target regulatory potential model available from NicheNet to identify potential ligands whose signalling converged on to the enriched TFs. Regulatory potential scores were derived by incorporating several data sources that covered ligand–receptor relationships, signal transduction and gene regulation events and then applying network propagation methods to quantify the signal flowing from a ligand through its receptors on to the signalling proteins, transcriptional regulators and, finally, target genes. For each TF in our list, we selected the top ten ligands with the strongest regulatory potential scores derived from the model. Only ligands that appeared at least five times in the top-ten list were selected for further analysis. Subsequently, we restricted the list of ligands to those whose receptors were expressed in more than 5% of IAF cells. Ultimately, this pipeline yielded 13 putative ligands, which were further expanded on the basis of ligand family membership.
Analysis of mouse spatial transcriptomics dataset
Tissues were processed with the cell segmentation staining kit; thus, counting of transcripts was not restricted to the nucleus. In total, a dataset of 2,792,913 cells × 480 genes was generated from 12 mouse large intestines, representing three biological replicates from the following conditions: Glis3f/f;cre treated with DSS, Glis3f/f;cre treated with water, Glis3f/f treated with DSS and Glis3f/f treated with water. Data were analysed using a similar workflow to that described above (see Supplementary Data 3 for the panel gene list used to define cell types).
In brief, cells with total RNA counts less than 10 were removed, followed by normalization and log transformation, regression of total RNA, and scaling. The top 40 principal components were used to construct the k-nearest neighbour graph, and Leiden clustering was applied (resolution = 0.5), followed by visualization with UMAP. Subsequent iterative rounds of clustering were performed by grouping the clusters into four main compartments: epithelial, fibroblast, stromal and immune, including lymphocytes and myeloid cells. The following markers were broadly used: Krt8, Epcam (epithelial cells); Pdgfra, Dpt (fibroblasts); Pecam1, Acta2, Des, S100b, Rgs5, Snap25, Ano1 (stromal cells); Cd79a, Mzb1, Cd3d, Trac, Gata3, Xcl1 (lymphocytes); and Tyrobp, Aif1, Ccr2, Fgr, C1qa, Cd68, Cd83, Trem2, Lyve1, Clec9a, Cd209a, Fpr1, Cxcr2 (myeloid cells).
Compartment-specific analysis and clustering was implemented using a similar workflow to that described above by setting Leiden algorithm parameter resolution = 1, except for the immune compartment, for which resolution = 1.5 was used. Post hoc inspection of clusters led to removal of poor-quality cells. Finally, for cell type annotation, we performed a Wilcoxon rank-sum test to define the markers specific to each cluster and inspected the spatial distribution of each cluster and the expression of cell-type-specific markers listed in Supplementary Data 3. Ultimately, we identified 55 unique cell types exhibiting distinct expression signatures and spatial distribution in the mouse large intestine.
Mice
Animal studies complied with all relevant ethical regulations. All mice used in this study were co-housed in specific-pathogen-free facilities at MGH. For all experiments, 5- to 20-week-old male or female sex-matched mice were used. Mouse strains included C57BL/6J Il11mNG mice, Il11f/f mice (C57BL/6JGpt-Il11em1Cflox/Gpt; GemPharmatech T051922), Glis3f/f mice (C57BL/6JGpt-Glis3em1Cflox/Gpt; GemPharmatech, T010949), R26-CreERT2 mice (B6.129-Gt(ROSA)26Sortm1(cre/ERT2)Tyj/J; Jax, 008463) and PDGFRa-CRE mice (C57BL/6-Tg(Pdgfra-cre)1Clc/J, Jax, catalogue no. 013148). No statistical calculations were performed to determine sample size. Genotype-matched mice were randomly assigned to each treatment group, and no blinding was performed. Mice were housed with a 12 h dark, 12 h light cycle, at an ambient temperature of 18–24 °C and 30–70% relative humidity, and provided with food and water ad libitum.
All animal procedures were conducted in accordance with protocol 2003N000158 approved by the MGH Institutional Animal Care and Use Committee, and animals were cared for according to the requirements of the National Research Council’s Guide for the Care and Use of Laboratory Animals.
Generation of Il11
mNG mice
Mouse Il11 was targeted with a gRNA verified in vitro (greater than 90% cleavage efficiency) near the stop codon of exon five (gRNA targeting sequence: TAAAGACTCGACTGTGACTC). Double-stranded DNA (dsDNA) was synthesized containing a T2A-mNeonGreen sequence flanked by approximately 400-nucleotide-long homology arms of the Il11 gene upstream and downstream of the stop codon. A single-stranded DNA (ssDNA) sequence of this donor template was synthesized with the Guide-it Long ssDNA Production System following the manufacturer’s protocol (Takara, catalogue no. 632644). Verified gRNA, ssDNA and Cas9 with NLS (PNA Bio, CP02) complex were microinjected into zygotes by the Harvard Transgenic Mouse Core. The resulting Il11mNG mice were backcrossed on the C57BL/6J background for three generations, followed by heterozygous crosses to obtain homozygous reporter mice. The licence to use mNeonGreen was acquired from Allele Biotechnology.
Generation of Il11
f/f;cre and Glis3
f/f;cre mice
Il11f/f mice (C57BL/6JGpt-Il11em1Cflox/Gpt; GemPharmatech, T051922) and Glis3f/f mice (C57BL/6JGpt-Glis3em1Cflox/Gpt; GemPharmatech, T010949) were generated by sperm reconstitution at the UMass Chan Transgenic Animal Modeling Core on behalf of GemPharmatech. Heterozygous pups were crossed together to generate homozygous conditional knockout mice. Homozygous Il11f/f mice were then bred with R26-CreERT2 mice (B6.129-Gt(ROSA)26Sortm1(cre/ERT2)Tyj/J; Jax, catalogue no. 008463) to generate homozygous Il11f/f, hemizygous cre mice. Glis3f/f mice were bred with hemizygous PDGFRa-CRE mice (C57BL/6-Tg(Pdgfra-cre)1Clc/J; Jax, catalogue no. 013148) to generate homozygous Glis3f/f, hemizygous PDGFRa-CRE mice.
Acute and chronic DSS colitis
DSS experiments were performed as previously described14. In brief, mice were given 2.0% (weight/volume) DSS (Thermo Fisher Scientific, J14489-22) dissolved in water for 7 days for the acute model of DSS. In the chronic model, DSS administration was followed by a water recovery phase lasting for another 7 days. This cycle was repeated two more times, with mouse weights recorded daily. Upon cessation of each model (at day 7 for the acute DSS model, day 42 for Il11 and Glis3 mouse experiments, or day 35 for cytokine neutralization experiments), mice were euthanized, and colons were obtained for phenotyping. For Il11 mice, chronic DSS experiments represented three pooled independent experiments with the following cohort replicates per experiment: Il11f/f, water (n = 3, 4, 11); Il11f/f;cre, water (n = 0, 0, 14); Il11f/f, DSS (n = 4, 3, 10); Il11f/f;cre, DSS (n = 3, 4, 3).
For Glis3 mice, chronic DSS experiments represented four pooled independent experiments with the following cohort replicates per experiment: Glis3f/f, water (n = 4, 4, 4, 6); Glis3f/f;cre, water (n = 0, 0, 0, 21); Glis3f/f, DSS (n = 4, 7, 3, 6); Glis3f/f;cre, DSS (n = 3, 4, 3, 5). For cytokine neutralization, chronic DSS experiments represented two pooled independent experiments with the following cohort replicates per experiment: n = 3, 4. Data pooling was performed by aligning animal data by timepoint (for instance, percentage weight change by day of DSS or water treatment), with all downstream analyses restricted to end point measurements.
Tamoxifen administration
Il11f/f and Il11f/f;cre mice were treated with DSS following the chronic DSS model. After cessation of each cycle of DSS administration, mice were intraperitoneally injected with tamoxifen (Sigma, T5648) dissolved in corn oil (Sigma, C8267) (200 µl of 20 mg ml−1). Tamoxifen was administered for 3 consecutive days, each at a different site of the mouse abdomen for each cycle of water treatment. Weights were monitored daily.
Cytokine neutralization
Il11mNG mice were treated with DSS following the chronic DSS model. At the start of each cycle of DSS administration, mice were intraperitoneally injected with monoclonal antibodies directed against IgG (BioXCell, BP0083), anti-TGFβ (BioXCell, BP0057), anti-IL-1β (Invivogen, mil1b-mab9-1T), or a combination of anti-TGFβ and anti-IL-1β. Antibodies were diluted in PBS and intraperitoneally injected at different sites of the mouse abdomen on the first and third days of each cycle of DSS administration (100 µl, 100 µg per mouse). Weights were monitored daily.
Mouse colon cryosectioning
Water- and DSS-treated mice were euthanized with CO2 in accordance with the Institutional Animal Care and Use Committee protocol. Colons were removed, flushed and rinsed in PBS (Sigma, D8537-500ml), then cut longitudinally to expose the lumen. Colons were rolled from the distal to the proximal end with forceps and placed in cryo-blocks (Tissue-Tek, catalogue no. 25608-916) containing OCT (Tissue-Tek, catalogue no. 25608-930) on dry ice. Frozen colon Swiss rolls were sectioned in a Leica CM1950 cryostat at −20 °C into 10 µm slices, which were placed on to glass slides (Fisherbrand, catalogue no. 22-230-900) for imaging; and 50 µm and 100 µm slices were collected for RNA and protein extraction, respectively.
Mouse colon staining
Immunofluorescence
Fresh-frozen tissue sections were fixed in 4% paraformaldehyde (Electron Microscopy Services, catalogue no. 15710-S) diluted in PBS (v/v) for 10 min at room temperature, followed by three washes in PBS. Tissues were then permeabilized in 0.2% Triton X-100 in PBS (v/v) for 10 min at room temperature, followed by three washes in PBS. Tissues were blocked with 4% bovine serum albumin (BSA; LGC Clinical Diagnostics, catalogue no. 1900-0016) in PBS (w/v) for 20 min at room temperature, then incubated with diluted primary antibodies in 4% BSA in PBS for 1 h at room temperature (1:400 for anti-mNeonGreen (Proteintech, #nfms), 1:400 for anti-CD68 (Cell Signaling Technology, 97778S)). Tissues were then washed three times in PBS and incubated with secondary antibodies in 4% BSA in PBS for 1 h at room temperature (1:500 AF-488 conjugated anti-mouse antibody (Proteintech, sms1AF488-1), 1:1,000 AF-594 conjugated anti-rabbit (Thermo Fisher Scientific, A-21207)).
Tissues were washed three times in PBS, rinsed in deionized water, mounted with ProLong Diamond Antifade Mountant with DAPI (Thermo Fisher, P36962) and sealed with a cover glass (Corning, catalogue no. 2975-245) for 24 h. Images were captured on a Nikon Ti2-E inverted microscope equipped with a CSU-W1 spinning disc confocal system.
QuPath v.0.6.0 was used to quantify the percentages of IL-11mNG fibroblasts in whole-colon Swiss rolls from Il11mNG mice subjected to chronic DSS. Channel minimum, maximum and gamma values were set to equal measurements across the sample cohort. Cell detection was performed on the basis of DAPI-stained nuclei. IL-11mNG cells were then defined by setting a single measurement classifier on objects filtered on all DAPI-stained cells. A channel filter was then set for ‘Cell: 488 nm mean’. An ‘Above Threshold’ was then set that defined IL-11mNG+ cells on the basis of the background signal distribution of 488 nm mean cell measurements in water-treated colon samples.
Masson’s trichrome staining
Staining was performed following the Masson’s Trichrome Stain Kit protocol (Vitroview, VB-3016). In brief, frozen tissue was acclimated to room temperature, fixed in 10% formalin for 30 min, and rinsed in distilled water three times. Tissues were then incubated in Bouin’s fluid for 1 h at 60 °C, washed in running tap water for 5 min, incubated with Weigert’s A+B solution mixture (1:1 mixture) for 7 min, and rinsed again with running tap water for 5 min, followed by incubation with Biebrich scarlet–acid fuschin stain for 5 min. Tissues were next rinsed in distilled water, then incubated with phosphomolybdic–phosphotungstic acid solution for a total of 10 min. Without rinsing, aniline blue solution was added for 15 min, followed by three washes in distilled water.
Tissues were briefly rinsed in acetic acid, dehydrated with 90% and then 100% ethanol for 2 min each, cleared in xylene (Sigma, 214736-1L) for 5 min, three times, and finally mounted with Cytoseal 60 (EMS, catalogue no. 18007) and sealed with a cover glass. Images were captured on a Nikon Ti2-E inverted microscope for detailed magnification images or a Leica Aperio VERSA scanning system for quantification.
For quantification of colonic collagen with Masson’s trichrome-stained colon samples, FIJI imaging quantification software v.1.54p was used on images captured on the Leica Aperio VERSA scanning system. Images were increased in brightness and contrast by setMinAndMax (−12, 228). Masson’s trichrome stain deconvolution was then performed using the Colour Deconvolution 2 plugin, setting the vectors to User Values defined by Colour(1)(R1: 0.412010759, G1: 0.849633569, B1: 0.329195889), Colour(2)(R2:0.587269268, G2:0.67744923, B2: 0.442919122), Colour(3)(R3: 0.804103811, G3: 0.584262903, B3:0.109790353). Only the blue collagen channel was used; red (muscle) and black (nuclei) signals were excluded from analyses. Collagen-positive pixel areas were identified by setting the signal threshold to 140 units to obtain the percentage of collagen-positive pixels relative to the total image area. To determine the tissue area, images were converted to 8-bit format and the signal threshold was set to 180 units; the area was then measured.
The final collagen percentage was obtained by dividing the calculated collagen-positive pixel area of the tissue by the total tissue area of the image.
H&E staining
Staining was performed following the Hematoxylin and Eosin Stain Kit protocol (Vitroview, VB-3000). In brief, frozen tissue slides were fixed in cold 80% methanol (Sigma, 179337-1L) at 4 °C for 5 min. The slides were then placed in room temperature PBS for 10 min, followed by a rinse in running deionized water for 2 min. Next, slides were placed in Mayer’s Hematoxylin Solution for 5 min and then rinsed in running tap water for 5 min; placed in 95% ethanol for 1 min and then eosin solution for 1 min 30 s; and dehydrated by being placed in two changes of 95% ethanol for 2 min each, and two changes of 100% ethanol for 2 min each. The tissues were cleared in xylene (Sigma, 214736-1L) for 5 min, three times, and finally mounted with Cytoseal 60 (EMS, catalogue no. 18007) and sealed with a cover glass. Images were captured on a Leica Aperio VERSA scanning system.
Histopathological quantification of tissue inflammation
Blinded histopathological assessment of H&E-stained tissues was carried out by a trained pathologist (A.R.S.). H&E-stained slides were graded by a standardized score of tissue injury and inflammation as described previously61. The total histological score ranged from 0 to 10 and was derived from the sum of four scoring criteria: mucosal ulceration, epithelial hyperplasia, lamina propria mononuclear infiltration and lamina propria neutrophil infiltration. Mucosal ulceration scoring criteria were either normal (0), surface epithelial inflammation (1), erosions (2) or ulcerations (3). Epithelial hyperplasia scoring criteria were either normal (0), mild (1), moderate (2) or pseudopolyps (3). Lamina propria mononuclear infiltration scoring criteria were either normal (0), slightly increased (1) or markedly increased (2). Lamina propria neutrophil infiltration were either normal (0), slightly increased (1) or markedly increased (2).
Hydroxyproline determination
First, 100-µm slices of OCT-embedded frozen colonic tissues were obtained by cryosectioning and flash frozen in liquid nitrogen. Tissues were crushed using manual dissociation on dry ice and incubated in Pierce RIPA buffer (Thermo Fisher, catalogue no. 89900) for 20 min. Following a freeze–thaw cycle, hydroxyproline extraction was performed following the manufacturer’s protocol with a few modifications (Sigma, MAK463-1KT). Fifty microlitres of lysates were incubated with 50 µl of 10 N NaOH for 2 h at 100 °C under constant agitation. Samples were allowed to cool to room temperature, then neutralized by addition of 50 µl of 10 N HCl, diluted 1:1 with 150 µl of water, and centrifuged for 5 min at 14,000g at room temperature to remove any particulates. Lysates were transferred to new tubes. A hydroxyproline standard (range: 0–50 µg ml−1) was then prepared, and 20 µl of each standard and tissue lysate were transferred to individual wells on a 96-well plate (Corning, catalogue no. 29444-018).
A reaction mixture containing 8 µl of reagent A and 90 µl of oxidation buffer was prepared for all samples, and 90 µl of this mixture was added to each well, followed by 10 min of incubation at room temperature. Then, 90 µl of reagent B was added to all wells, and the contents of the wells were mixed by pipetting up and down until turbidity dissipated. The plate was incubated for 90 min at 37 °C, after which the optical density of each well was measured at 560 nm on a BioTek Synergy H4 Hybrid Multi-Mode Microplate Reader. To quantify the amount of hydroxyproline, the optical density values of the standards were plotted against their concentrations. Each sample’s optical density measurement was then divided by the slope of the standard curve to yield a hydroxyproline concentration in micrograms per millilitre. Values were normalized to protein concentrations determined by measurements from a bicinchoninic acid (BCA) assay.
Protein concentration determination
Total protein concentrations were determined using a Pierce BCA protein assay kit (Thermo Fisher, catalogue no. 23225) on prepared tissue lysates with a few modifications. A standard curve with Pierce BSA standard (Thermo Fisher, catalogue no. 23209) was prepared (range: 0–10 µg µl−1) using individual wells of a 96-well plate, and 1 µl of sample was added to each well. A reaction mixture was then prepared by adding 50 parts of BCA reagent A with 1 part of BCA reagent B, and 200 µl of BCA reaction mixture was added to each well of the 96-well plate, followed by incubation of the plate for 30 min at 37 °C. The optical density of each well was measured at 562 nm on a BioTek Synergy H4 Hybrid Multi-Mode Microplate Reader. Each sample’s optical density measurement was then divided by slope of the standard curve to yield a protein concentration in micrograms per microlitre.
Mouse colon dissociation
Mouse colons were dissociated following a modified protocol62. Following acute or chronic DSS treatment, mice were euthanized, and colons were placed in ice-cold HBSS (Thermo Fisher, catalogue no. 14170112). Colons were cut longitudinally and then cut into small pieces. The tissue was then placed in 15 ml of epithelial cell solution (10 mM HEPES (Thermo Fisher, catalogue no. 15630080), 0.5 M EDTA (Thermo Fisher, catalogue no. AM9260G), 100 U ml−1 penicillin–streptomycin, 2% fetal bovine serum (FBS), 100 μg ml−1 DNase I (Roche, catalogue no. 10104159001) in HBSS). Samples were incubated at 37 °C for 15 min and manually shaken every 5 min. After incubation, samples were placed on ice for 10 min. Tissue pieces were then transferred to new tubes with HBSS and vigorously shaken for 40 s. The supernatant was collected by filtering through a 100-μm cell strainer (Corning, catalogue no. 352360) into a new tube, and pieces of tissue were placed in the original tube.
Shaking and filtering were performed a further two times to dissociate and deplete this epithelial fraction. The remaining tissue was placed in HBSS and washed with gentle shaking three times, for 30 s each time. With a razor blade, the tissue was cut into smaller pieces and transferred to a new tube containing 10 ml of lamina propria solution (100 U ml−1 penicillin–streptomycin, 100 μg ml−1 Liberase (Sigma, catalogue no. 5401127001), 100 μg ml−1 DNase I in RPMI). Tissues were incubated 37 °C for 20 min. Manual pipetting with a 5-ml pipette was used to break apart the tissue. The tissue was then incubated at 37 °C for a further 10 min, followed by manual pipetting with a 1-ml pipette to break apart the tissue further until dissociation. Samples were transferred to ice, and 1 ml of FBS and 80 μl of EDTA were added to stop the digestion reaction.
Tissue samples were pipetted again with a 1-ml pipette and filtered with a 40-μm cell strainer (Corning, catalogue no. 352340). HBSS was added, and cells were spun down at 400g for 10 min.
Cell surface staining of colonic cells
Isolated colonic cells were washed in PBS and resuspended in PBS with 2% FBS. Cells were then Fc blocked (BioLegend, catalogue no. 101320) on ice for 10 min at 1 μg per 100 ml buffer. Without removal of the Fc block, a master-mix of fluorophore-conjugated antibodies diluted 1:200 each in PBS with 2% FBS was added to the cells for 30 min on ice (APC anti-mouse CD45 (BioLegend, catalogue no. 157605), PE/Dazzle 594 anti-mouse TER-119 (BioLegend, catalogue no. 116243), Brilliant Violet 421 anti-mouse PDGFRA (BioLegend, catalogue no. 135923), PE anti-mouse PECAM1 (BioLegend, catalogue no. 102407), Brilliant Violet 711 anti-mouse EPCAM1 (BioLegend, catalogue no. 118233) and LIVE/DEAD Fixable Near IR (780) viability stain (Thermo Fisher, L34992)). Cells were spun at 500g for 3 min and washed with PBS. This spin and wash was repeated; then, cells were resuspended in live cell sorting buffer (PBS with 5% FBS, 25 mM HEPES, 1 mM EDTA) and sorted on a Sony SH800 cell sorter.
Single-antibody stains for each fluorescent channel using compensation beads (Beckman Coulter, B22804) were used to set up compensation.
Cell culture
Primary colon fibroblasts (CRL-1459), and immortalized fibroblasts (CRL-4001) were obtained from the American Type Culture Collection. Lenti-X 293T cells for lentivirus production were obtained from Takara (catalogue no. 632180). Primary-monocyte-derived macrophages were from isolated from healthy donor blood (unpurified buffy coats, 25–50 ml; ages within 18–65 years, unknown distribution of male and female) were collected at Research Blood Components, LCC, MA, USA, after a signed consent form had been obtained. Standard testing for blood-borne pathogens was performed. THP-1 Cas9-expressing cells were a gift from the Genomics Platform of the Broad Institute of MIT and Harvard. Commercially purchased cell lines were authenticated by the American Type Culture Collection with short tandem repeat identification. No authentication of Lenti-X 293T cells was provided by Takara, nor were donor or donated cells authenticated. All cell lines tested negative for mycoplasma except donor-derived cells, which were not tested.
Fibroblasts and Lenti-X 293T cells were cultured in Dulbecco’s modified Eagle’s medium (Thermo Fisher Scientific, catalogue no. 10569044) supplemented with 10% (v/v) heat-inactivated FBS (MilliporeSigma, F2442), 1× penicillin–streptomycin (Corning, 30-002-CI). Myeloid cells were cultured in RPMI-1640 medium (Gibco, catalogue no. 22400-089) supplemented with 10% FBS and 1× penicillin–streptomycin. All cells were cultured at 37 °C, with atmospheric O2 and 5% CO2.
Human monocyte purification and macrophage differentiation
Human monocytes from unpurified buffy coats were isolated with RosetteSep Human Monocyte Enrichment Cocktail (STEMCELL, catalogue no. 15028) following the manufacturer’s protocol, then enriched with SepMate tubes and Ficoll-mediated separation. Isolated cells were washed with PBS before being plated in RPMI-1640 containing recombinant M-CSF (PeproTech, catalogue no. 300-25) for 7 days on non-treated petri dish plates (Falcon, catalogue no. 351029). For polarization to different macrophage subsets, on day 6 following monocyte isolation, attached macrophages were washed with PBS and incubated with macrophage-polarizing ligands for 24 h (see Supplementary Data 4 for concentrations and catalogue numbers). For differentiation of THP-1 Cas9 monocytes into adherent macrophages, cells were stimulated with 300 ng ml−1 phorbol myristate acetate (Invivogen, tlrl-pma) for 3 h at 37 °C on non-treated petri dishes. Cells were then washed in PBS and reseeded on tissue-culture-treated plates for 2 days in RPMI-1640.
Cell stimulation
Fibroblasts were first seeded on a multiwell tissue-culture plate. Following adherence at 75–90% confluency overnight, cells were washed with PBS, and the appropriate ligand diluted in complete DMEM was added for the indicated time (Supplementary Data 4). Following stimulation, plates were spun down, and media supernatant was collected for analysis, or cells were washed with PBS and collected into TRIzol reagent (Thermo Fisher Scientific, catalogue no. 15596026) for RNA extraction.
Calculation of additive or synergistic IL-11 production
The following mathematical formulation published by Sanford et al.63 was used to determine whether IL-11 was produced additively or synergistically with dual TGFβ and IL-1β stimulation. The amount of IL-11 produced in response to single cytokine stimulation or at baseline was set as the following variables.
$$begin{array}{l}mathrm{Expression; of; IL}text{-}mathrm{11; at; baseline}={X}_{mathrm{baseline}} mathrm{Expression; of; IL}text{-}{rm{11; after; TGF}}beta ={X}_{mathrm{baseline}}+Delta {rm{TGF}}beta {rm{Expression; of; IL}}text{-}{rm{11; after; IL}}text{-}1beta ={X}_{mathrm{baseline}}+Delta {rm{IL}}text{-}1beta end{array}$$
$$begin{array}{l}{rm{I}}{rm{f}},{rm{e}}{rm{x}}{rm{p}}{rm{r}}{rm{e}}{rm{s}}{rm{s}}{rm{i}}{rm{o}}{rm{n}},{rm{o}}{rm{f}},{rm{I}}{rm{L}}text{-}11,{rm{a}}{rm{f}}{rm{t}}{rm{e}}{rm{r}},{rm{T}}{rm{G}}{rm{F}}{rm{beta }},{rm{a}}{rm{n}}{rm{d}},{rm{I}}{rm{L}}text{-}1{rm{beta }},{rm{i}}{rm{s}},{rm{a}}{rm{d}}{rm{d}}{rm{i}}{rm{t}}{rm{i}}{rm{v}}{rm{e}} ,,,=,{{rm{X}}}_{{rm{b}}{rm{a}}{rm{s}}{rm{e}}{rm{l}}{rm{i}}{rm{n}}{rm{e}}}+Delta {rm{T}}{rm{G}}{rm{F}}{rm{beta }}+Delta {rm{I}}{rm{L}}text{-}1{rm{beta }} ,,,=,388.8end{array}$$
$$begin{array}{c}mathrm{If; expression; of; IL}text{-}{rm{11; after; TGFbeta ; and; IL}}text{-}1beta ,mathrm{is; synergistic} =,{{rm{X}}}_{mathrm{baseline}}times mathrm{fold; change},Delta mathrm{TGF}{rm{beta }}times mathrm{fold; change},Delta mathrm{IL}text{-}1{rm{beta }} =,{{rm{X}}}_{mathrm{baseline}}times (({{rm{X}}}_{mathrm{baseline}}+Delta mathrm{TGFbeta })/{{rm{X}}}_{mathrm{baseline}})times (({{rm{X}}}_{mathrm{baseline}}+Delta mathrm{IL}text{-}1{rm{beta }})/{{rm{X}}}_{mathrm{baseline}}) =,{{rm{X}}}_{mathrm{baseline}}+Delta mathrm{TGFbeta }+Delta mathrm{IL}text{-}1{rm{beta }}+(Delta mathrm{TGFbeta }times Delta mathrm{IL}text{-}1{rm{beta }})/{{rm{X}}}_{mathrm{baseline}} =,1123end{array}$$
RNA isolation and RT–qPCR
In vitro cell cultures
RNA from in vitro cell cultures was isolated as previously described64. RNA was extracted from cells in TRIzol reagent following the manufacturer’s protocol with chloroform and isopropanol precipitation, followed by a wash in 75% ethanol (Thermo Fisher Scientific).
Tissue samples
RNA from in vivo tissue samples were isolated with an Aurum Total RNA Mini Kit (Bio-Rad, catalogue no. 7326820) following the manufacturer’s instructions. Then, 50-µm slices of OCT-embedded frozen colonic tissues were obtained by cryosectioning and flash frozen in liquid nitrogen. Tissues were crushed using manual dissociation on dry ice and then lysed in 700 μl of lysis solution, with pipetting up and down to lyse the tissue. The lysate was centrifuged for 3 min at room temperature at 14,000g and then transferred to a new tube; then, 700 μl of 60% ethanol was added to the lysate, and the mixture was transferred to an RNA binding column. The column was centrifuged for 60 s at room temperature at 14,000g, after which the filtrate was discarded. Next, 700 μl of low stringency wash solution was added to the column, and samples were centrifuged for 30 s at room temperature at 14,000g to remove the solution.
An 80-μl mixture of DNase I was added to the column and allowed to digest at room temperature for 25 min; after this, 700 μl of high stringency wash solution was added to the column, and samples were centrifuged for 30 s at room temperature at 14,000g to remove the solution. Low stringency wash solution (700 μl) was again added to the column, and samples were centrifuged for 30 s at room temperature at 14,000g to remove the solution. The column was then centrifuged for 2 min at room temperature at 14,000g to remove any remaining liquids. RNA was eluted by addition of 40 μl RNA elution solution, followed by centrifugation for 2 min at room temperature at 14,000g to collect the RNA.
Equal amounts of RNA were used to synthesize cDNA with an iScript cDNA synthesis kit (Bio-Rad Laboratories, catalogue no. 1708891). iTaq Universal SYBR Green Supermix (Bio-Rad Laboratories 1725124) was used for quantitative PCR with reverse transcription (RT–qPCR) on a C1000 Touch Thermal Cycler (Bio-Rad Laboratories). Relative gene expression was calculated using the 2−ΔΔCT method65, with HPRT as the reference housekeeping gene for human samples and Eef2 for mouse samples. See Supplementary Data 5 for primer sequences. Heatmaps of gene expression z-scores were generated in Microsoft Excel v.16.
Secreted protein quantification
Secreted cytokines were measured by spinning down cells and debris of cell culture media. IL-11, IL-1β and TGFβ were measured with a cytometric bead array assay (BD Biosciences, catalogue nos 560228, 558279, 560429) following the manufacturer’s protocol. Samples were measured for median fluorescence intensity of the cytokine of interest using a CytoFLEX S cytometer (Beckman Coulter). Median fluorescence intensity readings for each cytokine were plotted against a standard curve to obtain final cytokine concentrations. Data were analysed using FlowJo v.10.8 (BD Life Sciences)66.
Generation of CRISPR knock-in fibroblasts
IL11
mNG
Human IL11 was targeted with a gRNA verified in vitro (greater than 90% cleavage efficiency) near the stop codon of exon 5 (gRNA targeting sequence: TGAAGACTCGGCTGTGACCC). dsDNA was synthesized containing a T2A-mNeonGreen sequence flanked by approximately 400-nucleotide-long homology arms of the IL11 gene upstream and downstream of the stop codon. ssDNA was then synthesized with the Guide-it Long ssDNA Production System following the manufacturer’s protocol. Cells were electroporated with the ssDNA, along with Cas9 with nuclear localization signal (PNA Bio, CP02) and IL11-targeting gRNA using a Lonza 4D-Nucleofector X Unit kit (Lonza, AAF-1003X) with SE Cell Line 4D-Nucleofector X Kit S (Lonza, V4XC-1032). Cells were expanded, and single-cell clones were sorted for mNeonGreen expression following fibroblast activation.
Positive clones were then stably infected with lentivirus encoding Cas9 (pXPR_101) or dCas9–VP64 (pXPR_109) (plasmids were obtained from the Genetic Perturbation Platform of Broad Institute of MIT and Harvard), and activity for CRISPRko or CRISPRa was verified with gRNAs. Cells were passaged under blasticidin S HCl (Thermo Fisher Scientific, A1113903) to maintain Cas9 expression.
GLIS33xFlag
For 3xFlag-tagged GLIS3 fibroblasts, a 3xFlag sequence flanked by short homology arms to the GLIS3 insertion site was synthesized and electroporated (as described above for IL11mNG) into IL11mNG Cas9 fibroblasts, along with a gRNA targeting GLIS3 (gRNA targeting sequence: CCAAGAGAGCTTTTAGCCTT). Cells were single-cell cloned, and 3xFlag knock-in cells were selected and verified for Flag expression with an anti-Flag antibody (MilliporeSigma, F1804-200UG) in an image-based assay (‘Immunofluorescence staining’).
CRISPRko, CRISPRa and siRNA cell line generation
Lentiviral gRNA cloning
Oligos targeting a gene of interest were designed on the basis of CRISPick of the Genetic Perturbation Platform of the Broad Institute of MIT and Harvard. See Supplementary Data 5 for sequences. Oligos were annealed and phosphorylated using T4 ligation buffer (New England Biolabs, M0202L) and T4 PNK (New England Biolabs, M0201L). Esp3i-digested (Thermo Fisher Scientific, FD0454) pXPR_003 or pXPR_502 for CRISPRko or CRISPRa, respectively, were then ligated with annealed oligos using quick ligase (New England Biolabs, M2200L). Plasmids were transformed into One Shot Stbl3 chemically competent Escherichia coli cells (Thermo Fisher Scientific, C737303), single colonies were selected, and DNA was extracted using a spin miniprep kit (Qiagen, catalogue no. 27106). Correct integration of gRNA sequences was verified by Sanger sequencing.
Lentivirus production
Lenti-X 293 cells were seeded and transfected with a combination of psPAX2 (Addgene, catalogue no. 12260), VSV-G (Addgene catalogue no. 8454), and CRISPRko or CRISPRa vectors in Opti-MEM I reduced serum media (Thermo Fisher catalogue no. 31985088) and complexed to Lipofectamine 2000 (Thermo Fisher Scientific, catalogue no. 11668019). After 16 h of incubation of cells with the multiplasmid complex, media were replenished, and cells were allowed to grow for 48 h. Media were then collected and spun to remove debris, and the supernatant containing lentivirus was frozen at −80 °C. IL11mNG Cas9 or dCas9–VP64 fibroblasts were stably infected with lentivirus-containing gRNAs targeting the gene of interest. Cells were then selected using puromycin (Thermo Fisher Scientific, A11138-03) and blasticidin.
Ribonucleoprotein knockout
Knockout cells were generated with gRNA–Cas9 electroporation, following the recommendations of the manufacturer (Integrated DNA Technologies). In brief, a synthesized gRNA for the gene of interest was obtained through Integrated DNA Technologies and resuspended in water. The ribonucleoprotein complex was formed by incubating 120 pmol gRNA with PBS and 104 pmol of Alt-R Cas9 enzyme (Integrated DNA Technologies, catalogue no. 1081058) at room temperature for 20 min. Cells were electroporated with the resultant ribonucleoprotein complex using a Lonza 4D-Nucleofector X Unit kit with SE Cell Line 4D-Nucleofector X Kit S.
Small interfering RNA
Predesigned small interfering RNA (siRNA) oligomers were purchased from Sigma-Aldrich and resuspended in nuclease-free water at 20 μM (scrambled control (Sigma, SIC001), FRA1 (Sigma, SASI_Hs01_00191186), TEAD1 (Sigma, SASI_Hs01_00031302), TEAD3 (Sigma, SASI_Hs01_00090158)). Seeded fibroblasts were transfected with 150 pmol siRNA complexed with Lipofectamine RNAiMAX (Thermo Fisher, catalogue no. 13778030) in Opti-MEM media (Thermo Fisher, catalogue no. 31985062). Twenty-four hours later, cells were reseeded into 12-well plates and plated in triplicate knockdown per condition; another 24 h later, cells were washed with PBS and replenished with fresh media with or without addition of 10 ng ml−1 of human TGFβ and IL-1β for 24 h. Cells were washed in PBS and resuspended in TRIzol reagent for RNA isolation.
Fibroblast and macrophage co-culture
On day 6 after M-CSF treatment of primary human monocytes, differentiated macrophages were polarized with agonists for 24 h (for concentrations, see Supplementary Data 4). Following removal of agonist and several washes in PBS, fibroblasts were seeded with the macrophages for 24 h. For experiments using THP-1 macrophages, following 2 days of rest after phorbol myristate acetate stimulation, fibroblasts were seeded on to the differentiated THP-1 macrophages, and agonist was added to the media.
For qPCR-based measurements of fibroblasts in co-culture, fibroblasts were seeded at the bottom of a 24-well plate, with macrophages separated through a transwell insert (Corning, catalogue no. 3462). Following co-culture, macrophages were removed, and RNA was isolated from fibroblasts as described above.
Immunofluorescence staining
Monoculture of fibroblasts
Cells were seeded on a coverglass (Celltreat, catalogue no. 229173) in a 12-well tissue-culture dish. After completion of the experiment, they were fixed in 2% paraformaldehyde, followed by three washes in PBS for 5 min each time. Cells were then permeabilized with 0.2% Triton X-100, washed in PBS for 5 min, blocked with 4% BSA–PBS and incubated with 1:500 Flag antibody (MilliporeSigma, F1804-200UG) or 1:500 collagen 6 antibody (Thermo Fisher Scientific, PA5-106556) in 4% BSA–PBS for 1 h at room temperature. Cells were then washed again with PBS and incubated with a 1:1,000 dilution of AF-647-conjugated anti-mouse antibody (Thermo Fisher Scientific, A-21235) or a 1:1,000 dilution of AF-594 anti-rabbit antibody (Thermo Fisher Scientific, A-21207) in 4% BSA–PBS for 1 h. Finally, cells were washed three times in PBS, rinsed in distilled water, and mounted with ProLong Diamond Antifade Mountant with DAPI on to a glass slide for 24 h.
Images were captured on a Nikon Ti2-E inverted microscope equipped with a CSU-W1 spinning disc confocal system.
Co-cultures of fibroblasts and macrophages
Fibroblasts and THP-1 macrophages were prestained with fluorescent labelling dyes before co-culture. In brief, differentiated macrophages were labelled with CellTracker Orange CMRA Dye at 10 µM (Thermo Fisher Scientific, C34551) for 30 min at 37 °C in serum-free RPMI-1640 medium. Cells were then washed in PBS and seeded on to a coverglass in a 12-well dish in complete RPMI-1640 overnight. The following day, adherent fibroblasts were labelled with CellTracker Blue CMAC Dye at 10 µM (Thermo Fisher Scientific, C2110) for 30 min at 37 °C in serum-free RPMI-1640 medium. Cells were then washed, trypsinized and seeded on to the labelled macrophages along with macrophage-activating ligand FSL-1 (1 ng ml−1; Invivogen, tlrl-fsl) overnight. Immunofluorescent staining was performed as above (‘Monoculture of fibroblasts’). Images were captured on a Nikon Ti2-E inverted microscope equipped with a CSU-W1 spinning disc confocal. Cells were then washed with PBS and imaged again.
FIJI imaging quantification software (v.1.54p) was used to quantify the fluorescence intensity of AF-647-labelled nuclear GLIS33xFlag or AF-594-labelled collagen 6. The median fluorescence intensity of AF-647-labelled nuclear GLIS33xFlag was quantified in each individual fibroblast nucleus.
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism v.10. One-way analysis of variance (ANOVA) was used for comparisons among three or more groups. Two-way ANOVA was used to compare factorial groups. For Tukey’s, Dunnet’s and Dunn’s multiple-comparison tests, the family-wise error rate was set to a level of 0.05 (95% confidence interval). Multiplicity-adjusted P values less than 0.05 were considered to indicate statistical significance. All experiments were repeated at least twice as successful, independent experiments.
CRISPR screens
Lentivirus stocks of the Brunello CRISPRko and Calabrese CRISPRa libraries were obtained from the Genomics Platform of the Broad Institute of MIT and Harvard. Viral titres were calculated using a puromycin selection assay in which a target multiplicity of infection of 0.3 was selected so that each individual fibroblast could be targeted with one gRNA. IL11mNG Cas9 or IL11mNG Cas9–VP64 fibroblasts were infected with the determined titres of the CRISPRko or CRISPRa libraries, respectively, at 500× representation of each library for 24 h in DMEM supplemented with 10 µg ml−1 polybrene (MilliporeSigma, TR-1003-G). Fibroblasts were washed in PBS the following day, then incubated with 1 μg ml−1 puromycin and 3 μg ml−1 blasticidin in DMEM for 3 days. Following selection, fibroblasts were reseeded and expanded under selection.
Expanded fibroblasts were then reseeded in DMEM with no selection for 24 h; following attachment, they were washed with PBS and stimulated with a dual combination of TGFβ and IL-1β (10 ng ml−1 each) in DMEM for 24 h. After stimulation, fibroblasts were detached with trypsin, washed in PBS and resuspended in live cell sorting buffer (PBS with 5% FBS, 25 mM HEPES, 1 mM EDTA), and the top and bottom 15% of IL11mNG fibroblasts were sorted on a Sony SH800 cell sorter (see Extended Data Fig. 6a for the gating strategy). Collected fibroblasts were then spun down and frozen for subsequent DNA isolation and sequencing.
CRISPR gDNA isolation and sequencing
Fibroblast DNA was isolated with a QIAamp DNA Micro Kit (Qiagen, catalogue no. 56304) following the manufacturer’s instructions. Isolated DNA was then eluted in water, and PCR was performed as previously described with a P5 primer and a unique P7 primer for each sample67. Solid-phase reversible immobilization cleanup was performed with AMPure XP beads (Beckman Coulter, A63881), and concentrations were quantified with Qubit dsDNA HS assay (Thermo Fisher Scientific, Q32854). Each sample was run on a 2% E-Gel (Thermo Fisher Scientific, G402022), the dominant bands were excised, and 4 nM of pooled DNA was sequenced with an Illumina NextSeq 500 instrument (NextSeq 500/550 High Output v.2 kit, 75 cycles) with the following parameters: single end, index 8, read 51.
CRISPR screen analyses
PoolQ 2.2.0 (https://portals.broadinstitute.org/gpp/public/software/poolq) was used to map raw data to gRNA reads to obtain raw count data. EdgeR was used to identify differentially abundant (enriched) guides in sorted samples68, and STARS v.1.3 (https://portals.broadinstitute.org/gpp/public/software/stars) was used to rank the targeted genes on the basis of enrichment scores of multiple guides. Genes were selected if the perturbation was greater or equal to 3 and the fold change was greater than or equal to 2 with P < 0.05.
Bulk RNA-seq of stimulated fibroblasts
IL11mNG Cas9 and IL11mNG Cas9–VP64 fibroblasts were infected with non-targeting lentiviral gRNA or GLIS3 gRNA to make control, GLIS3 CRISPRko or CRISPRa fibroblasts. Fibroblasts were expanded under 1 μg ml−1 puromycin and 3 μg ml−1 blasticidin selection to select for a bulk genetically perturbed population. GLIS3-perturbed fibroblasts were then seeded in DMEM. The next day, cells were washed with PBS, and new medium with or without dual TGFβ and IL-1β (10 ng ml−1 each) was added at 0, 6, 12 and 24 h in a reverse time course stimulation. After 24 h, fibroblasts were washed with PBS, RNA was extracted with TRIzol, and each sample’s total RNA was quantified.
mRNA was purified with a Dynabeads mRNA direct purification kit (Thermo Fisher Scientific, catalogue no. 61012) with 40 ng of total RNA per sample. RNA-seq libraries were generated with previously described preparations69 and sequenced with a NextSeq 500/550 high output kit v2.5 (Illumina, 20024906).
Analysis of bulk RNA-seq of stimulated fibroblasts
Raw FASTQ reads were processed using FastQC70, Trim Galore (https://github.com/FelixKrueger/TrimGalore) and SortMeRNA71 to remove low-quality reads, adaptors and ribosomal RNA. Sequencing data were aligned to the human reference genome (GRCh38) using STAR, and gene abundance was quantified using Salmon72. Differentially expressed genes were identified using the DESeq2 R package73. Genes with P < 0.05 were considered to be differentially expressed.
scRNA-seq of stimulated fibroblasts and isolated colonic mouse fibroblasts from DSS treatment
scRNA-seq of TGFβ- and IL-1β-stimulated fibroblasts
Fibroblasts were stimulated as for bulk RNA-seq; however, upon completion of stimulation, at least 2,000 live single cells per sample were resuspended in PBS with 0.05% BSA and submitted for 10x Genomics single-cell sequencing (3′ transcriptome V3 with Cell Suspension using a HiSeq X system for up to 300 cycles).
scRNA-seq of sorted PDGFRA+ fibroblasts
Twenty-thousand live single cells resuspended in PBS with 0.4% BSA were submitted for 10x Genomics single-cell sequencing (Chromium Next GEM Chip G, v.3.1). Libraries were sequenced across 1.2 lanes of an Illumina NovaSeqX flow cell (10B reads, up to 300 cycles).
Analysis of scRNA-seq of stimulated fibroblasts
Gene expression matrices for each individual sample were obtained by aligning the FASTQ sequence reads to the reference hg38 human transcriptome using CellRanger v.3.1.0 (10x Genomics). Data were analysed using the scanpy package. Further, the count matrices of cell X UMI barcodes were filtered to exclude poor-quality cells, that is, those expressing fewer than 200 detected genes, more than 10,000 UMIs, or a mitochondrial gene fraction greater than 10%. Count matrices of each individual sample were merged and read-depth normalized using the standard logTP10K normalization procedure (the number of transcripts per 10,000 transcripts in a cell). The top 2,000 variable genes were then identified, and the normalized expression matrix was centred and regressed for effects of cell cycle scores (S phase and G2M phase) + UMI count per cell.
Dimensionality reduction was then performed on the top 60 principal components selected on the basis of inspection of the elbow plot, followed by the construction of a k-nearest neighbour graph and application of the Leiden clustering algorithm. The resulting clusters were visualized in the UMAP embedding space.
Analysis of scRNA-seq of isolated colonic mouse fibroblasts after DSS treatment
Gene expression matrices for six individual mice, three acute DSS-treated and three chronic DSS-treated, were obtained by aligning the FASTQ sequence reads to the reference mm10 mouse transcriptome using CellRanger v.7.0 (10x Genomics). One chronic DSS-treated sample failed the sequencing quality control metrics, demonstrating poor quality, and was excluded. Data were analysed using the scanpy package, following a workflow similar to that described in the ‘scRNA-seq analysis and cell type identification’ section. Altogether, we identified ten unique fibroblast cell types.
Analysis of mouse spatial transcriptomics dataset
A gene expression matrix of 1,540,301 cells × 480 genes was obtained from colons of three wild-type DSS-treated and three Glis3f/f;cre DSS-treated mice and analysed downstream using a similar workflow to that described in the ‘Analysis of human spatial transcriptomics dataset’ section. Altogether, we identified 41 unique cell types across the different compartments.
GLIS33xF
lag ChIP–seq
ChIP–seq was performed according to the protocol of Tian et al.74 with a few modifications. Following treatment of GLIS33xFlag fibroblasts with medium alone or TGFβ and IL-1β (10 ng ml−1 each) for 24 h, adherent fibroblasts were washed with PBS three times, then fixed with 0.25 M disuccinimidyl glutarate (Thermo Fisher Scientific, catalogue no. 20593). Fibroblasts were washed three times in PBS, then fixed with 1% formaldehyde (MilliporeSigma, catalogue no. 1040021000). Following three more washes, fibroblasts were scraped into PBS, spun down into pellets, and resuspended in SDS lysis buffer supplemented with protease inhibitor cocktail (Sigma-Aldrich, P8340-1ML). Lysates were frozen for at least 1 h at −80 °C, followed by thawing and sonication in a microTube-500 AFA fibre screw cap (Covaris, catalogue no. 520185) on a Covaris LE220Rsc Focused Ultrasonicator-110586. Sonication was performed at 7 °C, peak incident power 450, duty factor 30%, 200 cycles per burst, treatment time 100 s, and repeated four times.
Sonicated lysates were spun down and quantified for total protein and DNA using a BCA assay (Thermo Fisher Scientific, catalogue no. 23225), with measurement of absorbance at 260 nM (A260). Equal lysate concentrations were then brought up to 900 µl in low-ionic-strength chip dilution supplemented with 1 mM DTT (Thermo Fisher Scientific, R0861), and 4 µl of either IgG control (Thermo Fisher Scientific, 14-4714-82) or Flag antibody (MilliporeSigma, F1804-200UG) was added to the appropriate sample with rotation overnight at 4 °C. Following incubation, Pierce Protein G Magnetic Beads (Thermo Fisher, catalogue no. 88847) were added, followed by incubation for another 4 h at 4 °C. Beads were separated from the solution and washed twice with low-ionic-strength ChIP dilution buffer, once with high-salt buffer, once with LiCl, and finally twice with TE buffer following previously published methods74.
To elute from the beads, elution buffer was added for 15 min, beads were separated by magnet, and elution was repeated once more. Pooled eluates were then decross-linked overnight at constant rotation at 65 °C by addition of decross-linking buffer. Following incubation, 50 μg ml−1 RNase A (Qiagen, catalogue no. 19101) was added for 1 h at 65 °C; then, 0.4 mg ml−1 proteinase K (Thermo Fisher Scientific, AM2548) was added to digest proteins for 4 h at 65 °C. DNA was then extracted with phenol/chloroform/isoamyl alcohol (25:24:1, v/v) (Thermo Fisher Scientific, catalogue no. 15593031) extraction, and the DNA pellet was concentrated in a 2× volume of 100% ethanol, 1 μl of glycoblue coprecipitant (Thermo Fisher, AM9516) and 0.1× volume of sodium acetate (Thermo Fisher Scientific, AM9740). DNA pellets were spun down, washed in 75% ethanol, and resuspended in water. Samples then underwent library preparation for ChIP–seq using the KAPA HyperPrep Kit (Roche, catalogue no. 07962363001), following the manufacturer’s protocol.
Library yield was quantified using the dsDNA-specific assay Qubit (Thermo Fisher Scientific, Q33231). The size distribution of libraries was determined by Agilent 4200 TapeStation analysis (Agilent, G2991BA). Barcoded sample libraries at equimolar concentrations were pooled and paired-end sequenced using an Illumina NextSeq 2000 Sequencing System with NextSeq 2000 P3 Reagents Kit (200 cycles) (Illumina, catalogue no. 20040560), following the manufacturer’s instructions.
GLIS33xF
lag ChIP–seq analysis
ChIP–seq data were processed using the nf-core/chipseq 2.0.0 workflow75. Raw sequencing reads were preprocessed using FastQC and Trim Galore to remove low-quality reads and adaptors. The remaining reads were then aligned to a reference genome (GRCh38)76 using BWA. Picard’s MarkDuplicates (https://broadinstitute.github.io/picard/), SAMtools77 and BAMTools78 were used postalignment for filtering and removal of unmapped, multimapped, PCR duplicate and mismatched reads. BEDTools79 and bedGraphToBigWig80 were used to create normalized bigwig files, which were visualized using Integrated Genomics Viewer81. The retained high-quality alignment results were used to call narrow peaks using MACS2 (refs. 82,83) against an IgG control with q < 0.05. Consensus peaks set across all Flag antibody samples were created using BEDTools; featureCounts84 was used to count consensus peaks in each sample; and HOMER was used to annotate peaks relative to gene features and perform motif enrichment analysis.
Functional enrichment was analysed using the ClusterProfiler package85. Consensus annotated peaks enriched in 24-h samples were selected on the basis of having mean log2 fold change ≥ 0.15 relative to 0-h samples. Enriched peaks mapped to the nearest peaks were used for enrichment analysis according to the Gene Ontology Biological Processes database86. Pathways with Benjamini–Hochberg-adjusted P < 0.05 were considered for further analysis87.
ChIP–qPCR
Following treatment of fibroblasts with medium alone or TGFβ and IL-1β (10 ng ml−1 each) for 24 h, ChIP isolation was performed as described above (‘GLIS33xFlag ChIP–seq’) except that 4 µg each of ChIP-grade IgG control (Thermo Fisher Scientific, catalogue no. 14-4714-82), Flag antibody (MilliporeSigma, catalogue no. F1804-200UG), FOSL1 antibody (Cell Signaling, 5841S) or TEAD1 antibody (Active Motif, catalogue no. 61643) was used. Once the DNA pellets had been resuspended in water, the samples were subjected to qPCR with iTaq Universal SYBR Green Supermix for the indicated target genes (see Supplementary Data 5 for primers). Data were normalized using the fold enrichment method, in which IP signals are divided by the IgG signals, representing the IP signal as the fold increase in signal relative to the background. Heatmaps of gene expression z-scores were generated in Microsoft Excel v.16.
GLIS3 signature and PROTECT cohort analysis
Ordered probit regression, implemented using the polr function in the MASS R package88,89, was used to assess the association between GLIS3 signature gene expression and the clinical Mayo scores for each patient. Mayo scores were considered as the response ordinal variable, with control samples as the lowest category. The model was fitted using single-sample gene set enrichment (ssGSEA) scores and each gene’s expression independently as predictors. Bonferroni-adjusted P values of less than 0.05 were considered to indicate statistical significance.
Derivation of GSEA scores
ssGSEA scores were calculated from transcriptome profiles for each subject using the ssGSEA module (v.10.1.0) implemented in GenePattern90.
Deconvolution of bulk RNA-seq profiles
RNA-seq profiles for the PROTECT cohort were downloaded from GSE109142. Transcript per million data were deconvoluted to estimate the fraction of each cell type with the CIBERSORT algorithm implemented in R package RNAMagnet91. To define cell-type-specific markers, Wilcoxon rank-sum test was performed on the Integrated IBD Atlas. Only genes that had P < 1 × 10−8 and average log-scale fold change ≥ 0.75 compared to the background and were expressed in fewer than 20% of cells in the background population were considered further. Pseudobulk expression matrix of selected markers at cell-type-population level was used as input to CIBERSORT.
Human participants and ethical statement
The IRB of Mass General Brigham reviewed and approved the protocol, including a waiver of informed consent from the 16 patients recruited into the PRISM cohort at MGH. The study protocol complied with all relevant ethics guidelines and regulations. Excess tissues from clinically warranted surgical resections of patients with diverticulitis, CD and UC were collected for research purposes. All samples were deidentified.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
■ مصدر الخبر الأصلي
نشر لأول مرة على: www.nature.com
تاريخ النشر: 2026-01-07 02:00:00
الكاتب: Vladislav Pokatayev
تنويه من موقع “yalebnan.org”:
تم جلب هذا المحتوى بشكل آلي من المصدر:
www.nature.com
بتاريخ: 2026-01-07 02:00:00.
الآراء والمعلومات الواردة في هذا المقال لا تعبر بالضرورة عن رأي موقع “yalebnan.org”، والمسؤولية الكاملة تقع على عاتق المصدر الأصلي.
ملاحظة: قد يتم استخدام الترجمة الآلية في بعض الأحيان لتوفير هذا المحتوى.