سربت صفحة دعم Google للتو وصول Pixel Studio إلى Pixel 8

جوجل تطبيق بيكسل ستوديو، التي ظهرت لأول مرة مع سلسلة Pixel 9 العام الماضي، تشق طريقها أخيرًا إلى سلسلة Pixel 8 التي سيتم إصدارها عام 2023. التطبيق، الذي شق طريقه أيضًا إلى Pixel 10 افتراضيًا، أصبح الآن على خطى تطبيق Journal.
Pixel Studio: كل ما تحتاج لمعرفته حول النظام الأساسي لتوليد الصور بالذكاء الاصطناعي
البدء باستخدام Pixel Studio أمر بسيط
بالنسبة لأولئك غير المدركين، فإن التطبيق، الذي يتيح لك بشكل أساسي إنشاء أعمال فنية تم إنشاؤها بواسطة الذكاء الاصطناعي باستخدام المطالبات، يعمل أيضًا كمحرر، مما يتيح لك ضبط الصور الموجودة وتحسينها باستخدام المطالبات ذات الصلة. الآن، إلى جانب نقل التطبيق إلى هاتف Pixel 8، يبدو أن شركة جوجل تعمل أيضًا على طرح وظائف إضافية للتطبيق.
تم تسليط الضوء على التطورين لأول مرة من قبل الناس في 9to5Google بعد الاطلاع على التطبيق صفحة الدعم الرسمية.
الوصول إلى Pixel 8
وعلى الرغم من أن جوجل لم تصدر إعلانًا رسميًا عن التوسيع بعد، إلا أن صفحة دعم التطبيق، والتي تم تحديثها مؤخرًا، تشير إلى خلاف ذلك.
تقول صفحة الدعم: “في الوقت الحالي، يتوفر Pixel Studio فقط على Pixel 8 والإصدارات الأحدث”. في السابق، كانت الجملة نفسها تقول: “في الوقت الحالي، يتوفر Pixel Studio فقط على Pixel 9 والإصدارات الأحدث.”
تجدر الإشارة إلى أنه على الرغم مما تقوله الصفحة، فإن دعم Pixel 8 لا يتوفر مباشرة مع أحدث إصدار من Pixel Studio على متجر Play.
يحصل Pixel Studio على “صورة متحركة”
مرة أخرى، لم يتم الإعلان عن هذا رسميًا حتى الآن، ولكن يبدو أن تطبيق Pixel Studio يحصل على ميزة رسوم متحركة جديدة. وقد وصل هذا أيضًا إلى صفحة دعم Pixel Studio.
إنشاء الرسوم المتحركة
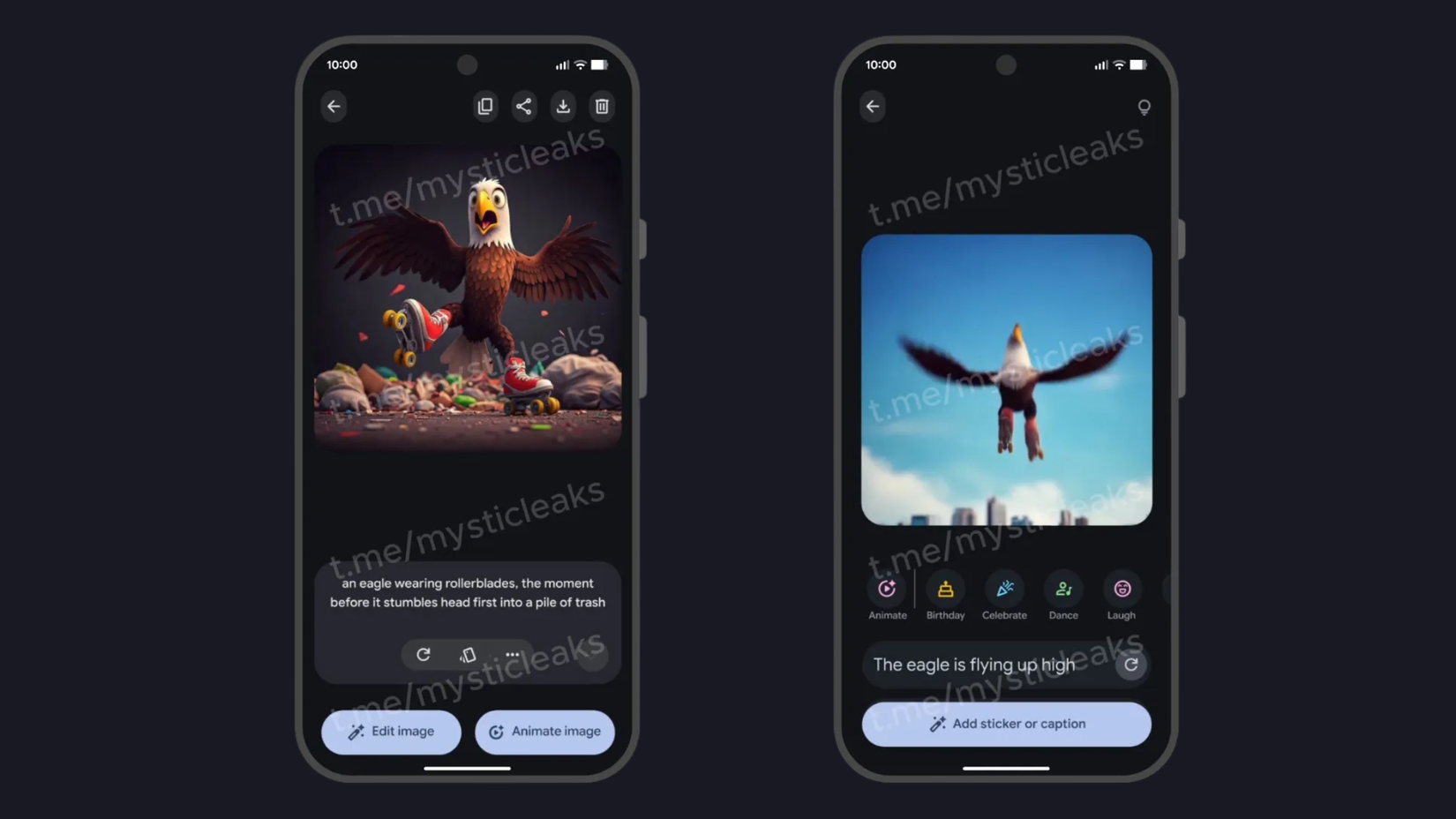
بمجرد إنشاء الصورة، يمكنك تحريكها.
- مقبض تحريك الصورة.
- أدخل مطالبة أو حدد مطالبة مقترحة أعلى مربع النص.
- بمجرد أن تصبح الرسوم المتحركة جاهزة، يمكنك إضافة ملصقات أو تعليقات عند النقر أضف ملصقًا أو تسمية توضيحية.
يمكنك بعد ذلك نسخ الرسوم المتحركة أو مشاركتها أو تنزيلها كملف GIF أو WebP.
ومن الجدير بالذكر أن هذه الميزة فعلت تسرب مرة أخرى في أكتوبر (من هنا الصورة أعلاه)، لكنها لم تصل إلى التطبيق مطلقًا. على الرغم من ذلك، فإن دعم الرسوم المتحركة أيضًا لا يتوفر مع أحدث إصدار من Pixel Studio، على الرغم من أنه بالنظر إلى الوثائق الموجودة على صفحة دعم عملاق التكنولوجيا، فإننا نعلم أنه سيأتي عاجلاً وليس آجلاً.
■ مصدر الخبر الأصلي
نشر لأول مرة على: www.androidpolice.com
تاريخ النشر: 2025-11-24 20:12:00
الكاتب: Karandeep Singh Oberoi
تنويه من موقع “yalebnan.org”:
تم جلب هذا المحتوى بشكل آلي من المصدر:
www.androidpolice.com
بتاريخ: 2025-11-24 20:12:00.
الآراء والمعلومات الواردة في هذا المقال لا تعبر بالضرورة عن رأي موقع “yalebnan.org”، والمسؤولية الكاملة تقع على عاتق المصدر الأصلي.
ملاحظة: قد يتم استخدام الترجمة الآلية في بعض الأحيان لتوفير هذا المحتوى.